Development of Omnitrap UVPD, ECD and EID LC-MS methods
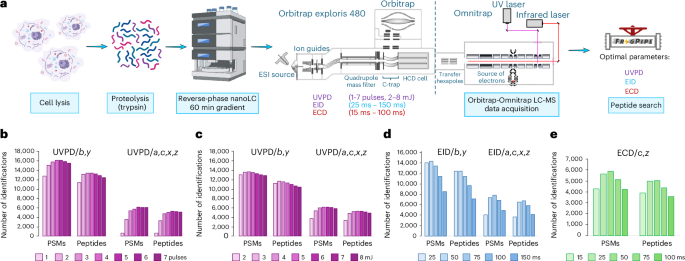
The results of our recent development and characterization of UVPD, EID and ECD on the Omnitrap platform36 suggested that it could be deployed in an LC-MS configuration for the analysis of complex peptide mixtures. Given that the conditions in direct-infusion experiments from our earlier work, such as number of available ions, injection times and ion transfer logistics, are typically more relaxed than in automated LC-MS analysis, an investigation is required to determine the optimal parameters for all dissociation techniques. Direct-infusion experiments reported previously36 were focused on higher resolution and signal-to-noise ratio with no regard to duty cycle. Given that acquisition of spectra with this configuration has limited parallelization potential (Extended Data Fig. 1a), we initially concentrated on reducing scan length to increase speed of spectra acquisition to handle the complexity of proteomes. The Omnitrap design requires ions to be cooled through a gas pulse prior to any ion manipulation. The original design used a single gas valve that had a maximum repetition rate of 10 Hz (Extended Data Fig. 1b). To improve the maximum rate of the Omnitrap we implemented the use of two valves, operating alternately, for gas injection, which can potentially double the speed (Extended Data Fig. 1b). Subsequently, we optimized the potentials for ion transfer in the Omnitrap to reduce the background collisional fragmentation (Supplementary Notes and Extended Data Fig. 1c–f). We then focused on increasing the identification rate in LC-MS experiments through application of pragmatic parameters for acquisition (Fig. 1a). Unless otherwise specified, human Expi293F cell lysate digests were used as the analyte. We began with the characterization of UVPD. We first varied the number of laser pulses at a fixed energy of 3 mJ per pulse and then varied the energy for a fixed number of pulses. For data analysis, we started with using only b and y ions for identification, which were previously shown to be the most abundant in UVPD of tryptic peptides6,37,38. Analysis shows that increasing the number of laser pulses leads to a greater number of identified peptide–spectrum matches (PSMs) and peptide sequences until a maximum is reached at four pulses (Fig. 1b). Further increases in the number of laser pulses used for dissociation results in a drop of the identification rate, either due to secondary fragmentation or reduced scan rate. We selected four pulses for further investigation and varied the energy of each pulse. In this series of experiments, the maximum of identified PSMs and peptide sequences was observed at distinct energies depending on the type of fragment ions used for identification (Fig. 1c). Using only b and y fragments, the maximum is observed at 5 mJ per pulse, while when other types of fragment characteristic of UVPD are used, namely a, c, x, z (ref. 4) (see Supplementary Table 1 for structures and definitions of fragment ions considered in this work), the maximum is located at 6 mJ per pulse. Given that a, c, x, z in contrast to b, y are more unique to UVPD, we opted to use 6 mJ per pulse in future experiments.
a, Experimental workflow. b–e, Number of PSMs and peptides identified in UVPD experiments varying the number of UV laser pulses at 3 mJ pulse−1 (b), UVPD experiments using four laser pulses and varying the pulse energy (c), EID experiments varying the irradiation time at 25 eV of electron energy (d), and ECD experiments varying the irradiation time at ~1 eV of electron energy (e). In UVPD and EID, b, y or a, c, x, z fragments were used for data analysis; c and z ions were used in the analysis of ECD data. Schematic diagram in a created in BioRender; Govender Kirkpatrick, M. https://biorender.com/qqloq0m (2025).
Next, we studied the optimal reaction times for ExD. In typical ExD experiments, ions are transferred into the reaction chamber and undergo irradiation by electrons emitted by a heated filament35 during a specified amount of time (Extended Data Fig. 1g,h). In EID experiments, we varied the irradiation time from 25 ms to 150 ms and measured the number of identified PSMs and peptides. We observed that b and y ions can be the most prominent ions in EID. When using only these two ions for analysis, the number of PSMs and of peptides reaches the maximum value at 50 ms of irradiation (Fig. 1d). At longer irradiation times, these numbers start to drop. Interestingly, the profile of peptide identification shows a much more distinctive dependence on the type of ions used for analysis compared with UVPD (Fig. 1d). At shorter irradiation times, a, c, x, z fragments are underrepresented compared with those of b, y, and the largest number of PSMs and peptides was observed at 75 ms (Fig. 1d). To keep scan rates high in the interest of absolute number of identifications, we chose to continue with the 50 ms irradiation time. Finally, we found 50 ms of irradiation to be optimal in ECD using c and z fragments for the data analysis (Fig. 1e). We did not investigate other main-series types of fragments, because the majority of the products of ECD of relatively short peptides are c and z ions7,8. Given that ECD is known to be a charge-dependent process favoring higher charge states, the value of 50 ms obtained using mainly doubly charged and less frequently triply charged precursors of tryptic peptides can be considered conservative. To characterize the fragmentation behavior of ECD, UVPD and EID, a larger and more diverse range of peptides is required.
Large-scale multi-enzyme LC-MS analysis
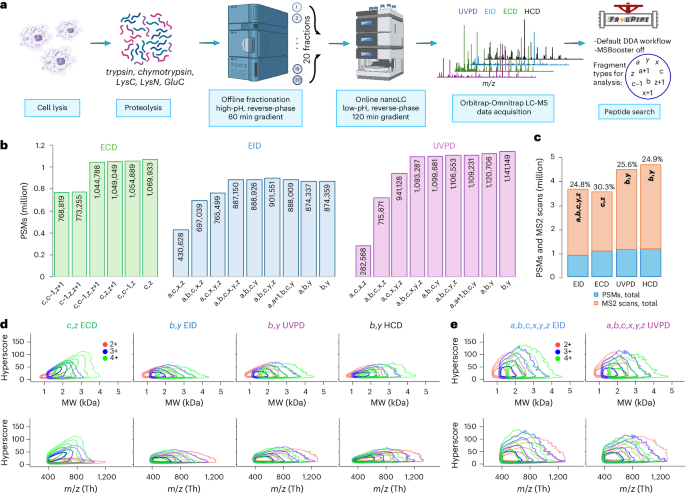
We increased the diversity of peptide sequences through the use of more proteases, and we increased peptide depth by utilizing offline reverse-phase high-pH fractionation (Fig. 2a). We chose trypsin, LysC, GluC, chymotrypsin and LysN because they have been shown to produce complementary results in terms of peptide length, protein sequence coverage, and frequencies and positions of amino acid residues across the peptide backbone39. Next, we fractionated each digest40 into 20 pooled fractions and analyzed all of them using ECD, EID, beam type CID (referred to as higher-energy CID or ‘HCD’ on Thermo instrumentation) and UVPD LC-MS. The choice of liquid chromatography gradient time for the dissociation techniques was based on their maximum sequencing rate to ensure that they all produced a similar number of scans.
a, Experimental workflow. b, Total numbers of PSMs in ECD, EID, and UVPD experiments identified using different combinations of fragment types. c, Highest number of PSMs from b (blue) and total number of acquired MS2 scans (orange) in ECD, EID, UVPD and HCD experiments; the rate of PSM identification is shown above each corresponding bar. d, Density contour plots of hyperscore distributions of 2+, 3+ and 4+ charge states of unique PSMs (unique combination of amino acid sequence, charge and modification selected by highest hyperscore) acquired in ECD using c and z fragments and in EID, UVPD and HCD using b and y fragments. e, Density contour plots of hyperscore distributions of 2+, 3+ and 4+ charge states of unique PSMs acquired in EID and UVPD using a,b,c,x,y,z fragments. Contour lines demarcate the smallest regions to contain 50%, 80%, 95% and 99% of points. Schematic diagram in a created in BioRender; Govender Kirkpatrick, M. https://biorender.com/4anifnk (2025).
The analysis of UVPD, EID and ECD data is not as straightforward as that of HCD data. The major products of HCD are well characterized, with a, b, y ions dominating data. In contrast, UVPD and EID are known to produce all main-series types of peptide fragments as well as some radical a + 1, x + 1 ions,4,15,41 with the last two largely understudied. The average proportion of each type of main-series fragment has been reported for UVPD6,37,38; however, the effects of using these ions and their combinations in the automated data analysis have not been extensively discussed. We therefore analyzed the acquired raw data using several unique combinations of the expected fragment types with the goal to maximize the number of identified PSMs while maintaining the same 1% false discovery rate (FDR). For ECD, the most important ions for robust identification were c and z (Fig. 2b). The addition of c − 1 or z + 1 had a minimal and slightly detrimental effect. Analogously, b and y were the dominant ion types for both EID and UVPD. However, a, a + 1, c, z ions were beneficial for improving identification rates for EID, while b, y produced the best results in UVPD. The numbers when broken down to the individual enzyme level are similar to the global result, although tryptic and LysC peptides enhance the formation of z + 1 ions while impairing the formation of c − 1 in ECD, and favor the generation of y ions in EID and UVPD compared with other enzymes (Supplementary Fig. S1). The results for UVPD and EID seem to be strongly dependent on y ions and to a smaller degree on b ions. While no extensive literature exists for EID, our UVPD data agree with previous findings. Others also found that b, y fragments are the most abundant types of ions in 193 nm UVPD of tryptic peptides, and the ion current of y fragments is approximately double that of b (refs. 6,37). Similarly, b, y fragments dominate the spectra in 213 nm UVPD of tryptic peptides, and the average number of annotated y fragments is twice that of b ions38.
In total, each fragmentation technique produced between approximately 3.5 million and 4.5 million MS2 spectra across five enzymes, 20 fractions per enzyme (Fig. 2c). EID data had the least number of PSMs ( ~900,000), while UVPD, which has the fastest acquisition rate among all Omnitrap techniques studied here ( ~6.3 MS2 scans per second on average), had 1,141,000 (Fig. 2c). Surprisingly, charge-dependent ECD came closest to UVPD with 1,070,000 PSMs, even though its scan rate ( ~5.2 MS2 spectra per second) was essentially the same as in EID. HCD showed the highest numbers with 1,160,000 PSMs acquired using 60 minute gradients at the rate of, on average, ~13 MS2 scans per second. Pleasingly, the efficiency of peptide sequencing by EID (24.8%) and UVPD (25.6%), expressed as the ratio of the number of confidently identified PSMs to that of acquired MS2 scans, is essentially the same as by HCD (24.9%), while the efficiency of sequencing by ECD (30.3%) was the best (Fig. 2c). This was surprising considering the relative inefficiency of ECD for doubly charged peptides, which represent a substantial subset of identified peptides (Extended Data Fig. 2a).
The MSFragger hyperscore can serve as an indirect measure of the number of fragments found in a spectrum, similar to a spectrum quality score42. We plotted density contour plots for hyperscores of all unique precursors (that is, unique combinations of amino acid sequences, charge states and modifications, Extended Data Fig. 2b,c) per charge state using c, z fragments in ECD and b, y fragments in UVPD, EID and HCD (Fig. 2d and Supplementary Figs. S2 and S3). Expectedly, the distribution of hyperscores in ECD is strongly charge dependent, with doubly charged precursors assigned substantially lower values. Furthermore, the hyperscore distributions for 3+ and 4+ precursors in ECD have an apparent maximum at 800 Th. A similar trend was reported earlier by Good et al. for ETD of tryptic and LysC peptides, in which the percent of bonds cleaved by ETD begins to drop at approximately 600 Th for 3+ precursors and 650 Th for 4+ ones13. When analyzing solely b, y ion series, EID, UVPD and HCD all produce very similar hyperscore distributions for the same charge states of precursors (Fig. 2d). UVPD has marginally higher hyperscores in the low-m/z range than HCD, and EID produces lower hyperscores in the high-m/z range than UVPD and HCD. The upper boundary of hyperscore distributions for these dissociation techniques starts to drop beyond approximately 2,000–2,500 Da for 2+ and 3+ precursors and 2,500–3,000 Da for 4+ precursors. We interpret these observations as the reduction of the signal-to-noise ratio that follows the spreading of available fragment signal across a larger number of produced fragments in spectra of long and highly charged peptides, that is, signal splitting. The difference in number of identifications with the same 1% FDR was marginal for UVPD and EID when we increased the number of fragment types all the way up to a, b, c, x, y, z, as long as the b, y fragments were included (Fig. 2b). We therefore investigated how the choice of type of fragment for analysis affects hyperscores (Fig. 2e and Supplementary Fig. S4). Clearly, adding more types of fragments results in greatly improved hyperscores for both EID and UVPD, indicating a larger number of dissociated bonds and data-rich spectra.
Deep learning modeling of UVPD, EID and ECD fragment intensities
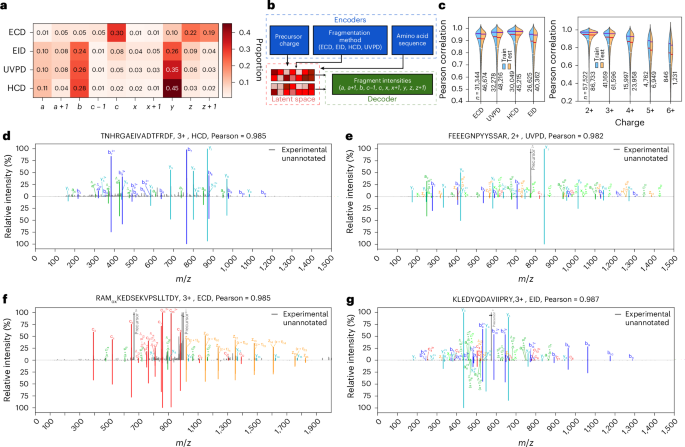
PSM scoring can be improved substantially if performed against experimental or in silico-generated spectral libraries32. Deep learning models have demonstrated promising results in predicting CID-based spectra of peptides using only peptide sequence, charge state and collision energy as input26,27,28,31, but no such models exist for other fragmentation techniques due to the lack of large amounts of high-quality data for training. We therefore set out to use the datasets generated in this work to train a deep learning model able to predict fragment ion intensities. To create a more comprehensive model we then generated a similar dataset for electron-transfer/collision-induced dissociation (ETciD) on a Thermo Tribrid instrument (Supplementary Notes). Training a deep model requires converting the raw data into a dataset containing correctly annotated peak intensities. This implies that we need to solve potential clashes such as, for example, a + 1 ion, which is a radical a ion coupled with an additional hydrogen atom, versus the 13C peak for an a ion. For all datasets, we performed an automated annotation of major fragment types expected in EID, ECD, ETciD and UVPD (Supplementary Table 1) using the Oktoberfest framework30. The comparison of [a + 1]/[a] ratio in HCD, EID and UVPD suggests that a large proportion of a + 1 in EID and UVPD spectra originate from gas-phase electron- and photon-based chemistries (Fig. 3a, Extended Data Fig. 3, Supplementary Figs. S5–S9 and Supplementary Notes). With the annotated spectra in hand, we defined our model’s ion dictionary and curated training and validation datasets. The original Prosit model27 architecture was designed around a structured output space consisting of b and y fragments with lengths 1–29 and charges +1 to +3. By contrast, the model trained on our data has an unstructured output space, with fragment ions chosen based on frequency of occurrence (≧100 occurrences, Supplementary Figs. S5–S9). The model also takes the categorical fragmentation type as input; given that the HCD data were acquired on a single instrument, it was unnecessary to use collision energy as additional input to the model, as was performed for previous Prosit models27. Our model shares similarity with the original Prosit model in that the sequence and metadata are separately encoded into latent spaces and combined in the interior of the network, but the metadata have slightly changed, and the model outputs predicted intensities of 815 fragment ions of various length, charge and fragment type (Fig. 3b). Results show very little overtraining: the median Pearson correlations for ECD, UVPD, HCD and EID are 0.919, 0.931, 0.950 and 0.897, respectively, on the training set, and the corresponding scores for the test set are only ~0.005 lower for each fragmentation method (Fig. 3c and Extended Data Fig. 4). Furthermore, we observe that precursor charge is consequential for prediction performance, with precursor charges greater than 2 having an increasingly wide range of Pearson correlations, likely to be due to the sparsity of high charge precursors in the training set and increasingly complex fragment ions present in the spectra. Pleasingly, we see that conditioned on the fragmentation method the model reliably assigns appreciable intensity only to those fragments expected for each fragmentation method, for example b, y for HCD and c, z for ECD (Fig. 3d,e). The model is also able to predict intensities of b, y and minor fragments, such as a, a + 1, x, x + 1, c, z in UVPD and EID, although predictions of low-intensity ions for the latter seem slightly less accurate (Fig. 3f,g). We performed a series of additional tests to validate the robustness and correctness of our model (Supplementary Notes and Supplementary Fig. S10).
a, Heatmap of mean proportion of each type of fragment ion among all annotated peaks in ECD, EID, HCD and UVPD spectra acquired across all enzymes, not reflecting relative intensity of ions. Annotation was performed for 10 ion types: a, a + 1, b, c − 1, c, x, x + 1, y, z, z + 1 (Supplementary Table 1). b, The modified Prosit deep learning architecture for prediction of fragment ion intensities in ECD, EID, HCD and UVPD spectra. The input parameters (peptide sequences, precursor charge state and fragmentation method) are encoded into a latent representation (latent space). This representation is then decoded to predict fragment ion intensities. c, Pearson correlation coefficients between predicted and experimental spectra in training and test sets separated by fragmentation method (left) and charge state (right). Horizontal white, red, and blue lines correspond to 25%, 50% and 75% percentiles, respectively. n indicates sample size. Distributions extending beyond 1.0 are plotting artefacts. d–g, Mirror plots of selected precursors in HCD (d), UVPD (e), ECD (f) and EID (g) data. Each mirror plot compares experimental (top) and predicted (bottom) fragment intensities, with each fragment type uniquely colored.
Rescoring of alternative fragmentation data using fragment intensity predictions
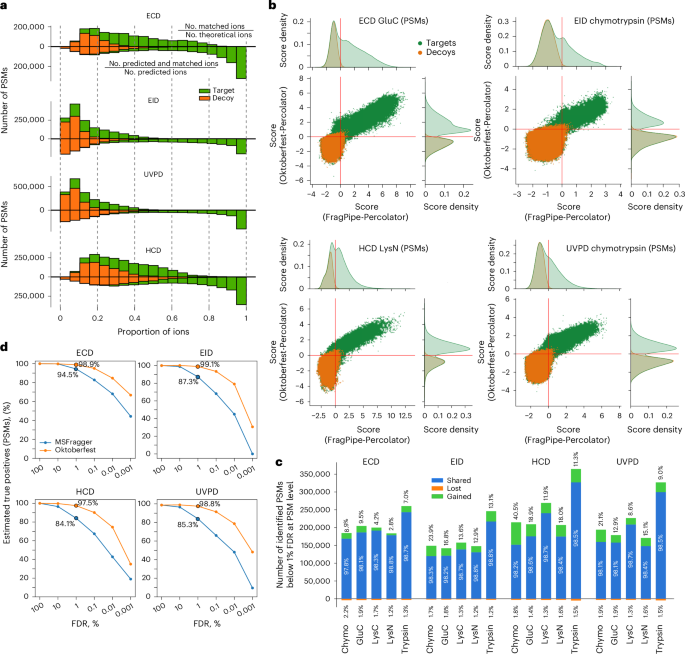
An efficient control of FDR in database searching is critical for identification of true-positive peptide matches. Previously, we showed that data-driven rescoring of CID data using the Prosit model greatly improved number and accuracy of peptide identifications27. We hypothesized that predicting fragment ion intensity would be beneficial for improving the results of the database searches of UVPD, EID and ECD data as well. Using the optimized MSFragger results we first calculated the ratio of the number of all observed to that of all possible theoretical fragment ions in each identified spectrum (Fig. 4a and Extended Data Fig. 5, upper distributions). The resulting distributions for target and decoy (a priori false-positive) PSMs were heavily intermixed and shifted towards smaller ratios. EID and UVPD ratios were particularly small due to a large number of theoretical ions. We then calculated the same ratios but allowed only fragments predicted by Prosit (Fig. 4a and Extended Data Fig. 5, lower distributions). The inclusion of only predicted fragments split the distribution of ratios of target PSMs, in which the majority shifted towards higher values with a larger portion being above 0.8, and the remainder were essentially unchanged. At the same time, the ratio of decoy PSMs remained clustered at lower values. This indicates a substantial improvement in the alignment between the observed and predicted fragment ions.
a, Histogram of the ratio of experimentally observed ions to all theoretically possible fragments (upper distributions); and histogram of the ratio of predicted and experimentally observed ions to all predicted ions (lower distributions). b, Correlation of Percolator scores for all target and decoy PSMs obtained from the rescoring of the MSFragger (top) and Oktoberfest (right) sets of scores for selected combinations of enzyme and fragmentation technique. The red solid lines indicate the 1% PSM-level FDR cut-offs. For database search scores, the best combinations of fragment types from Fig. 2b were used; for Oktoberfest scoring, most frequently annotated fragment types ( >4% of all annotated ions across all spectra) were used for each dissociation method (Extended Data Fig. 3). c, Number of shared, gained and lost PSMs identified at 1% PSM-level FDR using the Oktoberfest set of scores compared to the original MSFragger search for each fragmentation technique per enzyme. The numbers correspond to the data from b and Supplementary Figs. S11–S14. Chymo, chymotrypsin. d, Proportion of the number of true-positive PSMs to the estimated maximum number of true-positive PSMs acquired using original MSFragger and Oktoberfest scores at different values of PSM-level FDR for each fragmentation technique, all enzymes combined.
Next, we applied data-driven rescoring using the Oktoberfest framework, which benefits from the here-developed fragment ion intensity prediction model by generating fragment intensity-dependent scores rather than relying only on the presence or absence of any theoretical fragments. In combination with Percolator43, these scores are aggregated into a single score that maximizes the separation of correct and incorrect matches. The resulting Oktoberfest scores were then compared to the Percolator-derived scores from MSFragger (Fig. 4b and Supplementary Figs. S11–S15), which do not include fragment intensity-based features. For MSFragger database searches, we chose the best combination of ion types for each fragmentation method from Figure 2b, and for rescoring in Oktoberfest we used all of the most frequently annotated types of fragments ( >4% of annotated ions in a spectrum, averaged across all spectra) for each fragmentation technique (Extended Data Fig. 3). Both sets of scores were filtered to 1% FDR using Percolator43. While rescoring led to remarkable separation of decoys from targets for the majority of enzyme–fragmentation method pairs (Fig. 4b and Supplementary Figs. S11–S15), ECD in general demonstrated sufficient separation in database searches, such that rescoring delivers only marginal improvements in identification (Supplementary Fig. S11). This partly explains the highest identification rate observed for ECD in the initial database searches (Fig. 2c). We attribute this to the relative cleanliness of ECD spectra that consist primarily of c, z fragments, precursor ions and charge-reduced species, thus reducing chances for random false matches. Interestingly, ECD was the only technique in which it was possible to discriminate the distributions of charge states among target PSMs after rescoring, which reflects the distinct charge-dependent kinetics of this process (Supplementary Fig. S16). Using rescoring, we were able to salvage a substantial number of PSMs in all combinations of enzyme and dissociation method (quadrant II in Fig. 4b and Supplementary Figs. S11–S15). At the same time, a high number of PSMs initially identified were discarded (quadrant IV in Fig. 4b and Supplementary Figs. S11–S15).
To evaluate how this separation of scores translated into gains and losses of PSMs and peptides, we compared the results of the database search and rescoring at both 1% PSM-level (Fig. 4c) and 1% peptide-level FDR (Supplementary Figs. S17 and S18). The number of gained PSMs varied (depending on the enzyme and fragmentation method) between approximately 3% and 40.5%, with chymotrypsin HCD data producing a notable gain of 40.5%. The latter observation is consistent with our previous findings27. Remarkably, chymotrypsin was also the main beneficiary of rescoring in UVPD and EID data. This demonstrates the usefulness of rescoring for expanded search spaces characterized by an increased number of possible charge states, allowed missed cleavages and reduced enzyme specificity, all of which are typical for chymotrypsin (Extended Data Fig. 2a). Consistent with the score distributions (Fig. 4b and Supplementary Figs. S11–S15), ECD had the lowest number of gained PSMs and peptides regardless of protease among all fragmentation techniques (Fig. 4c and Supplementary Fig. S17). Further investigation of ECD data shows that prediction of retention time and of fragment intensity generated similar gains, each adding approximately 6.5% of PSMs (Supplementary Notes and Extended Data Fig. 6). Such a relatively modest contribution of retention time predictions shows that improvements observed after rescoring of other combinations of enzyme and fragmentation technique are primarily driven by the new Prosit model.
To explore the reasons for the varying number of gains observed, we investigated the recovery of estimated true-positive PSMs. We compared the number of estimated true positives across a range of FDR thresholds (by subtracting the number of decoy PSMs from the number of target PSMs at different FDR cut-offs) before and after rescoring with the total number of estimated true positives in the dataset that could be recovered from the initial search results, by subtracting the total number of decoys from the total number of target PSMs (Fig. 4d and Supplementary Fig. S19). At 1% PSM-level FDR, rescored ECD, EID and UVPD searches recovered more than 97% of possible true positives, while the original database searches extracted approximately 95% in ECD, 87% in EID, 85% in UVPD, and 84% in HCD. At a stricter FDR of 0.01%, the results after rescoring still captured more than 75% of all estimated possible true positives, with ECD showing the highest proportion approaching 85%. At the same FDR level, initial database searches identified less than 70% of possible true positives in ECD and less than 55% in all other dissociation methods (Fig. 4d). The analysis shows that data-driven rescoring using the pan-fragmentation Prosit model substantially increases the proportion of estimated true-positive PSMs retained at stringent thresholds, approaching saturation of the set of PSMs recoverable from the initial MSFragger search results. It is important to note that further correct identifications, for example from modified peptides not considered in the initial search, cannot be considered in the estimation of the number of true positives.
The rescoring data provided an opportunity to inspect the efficacy of each enzyme and dissociation technique for proteome analysis (Supplementary Notes, Extended Data Figs. 7 and 8 and Supplementary Figs. S20–S24). Trypsin, as expected, identified the most PSMs, peptides and proteins for every fragmentation technique. Chymotrypsin had the next best result, with LysC and LysN slightly further behind (Extended Data Fig. 7a and Supplementary Fig. S20a), replicating previous trends observed for CID and ETciD data44,45,46. The enzyme GluC clustered with LysN, appearing to be slightly superior or inferior depending on the dissociation technique. Average protein sequence coverage was similar for each fragmentation technique (Extended Data Fig. 8). To assess complementarity at the protein sequence level we represented our data at the amino acid level. In general terms, when comparing the complementarity of trypsin against its alternatives, we saw substantial improvements in proteome coverage for all fragmentation techniques (Extended Data Fig. 7b and Supplementary Fig. S20b); in fact, the unique combined coverage for LysN, LysC, GluC and chymotrypsin was more than that for trypsin. These observations echo previous work demonstrating the complementarity of enzymes for improving sequence coverage39,44,45,46. It should be noted that each trypsin fraction was essentially analyzed with LC-MS four times, and a more exhaustive LC-MS analysis would not significantly increase proteome coverage, and hence the amount of analysis time for the other enzymes versus trypsin is not an important factor in the comparison. Further analysis of unique coverage for each fragmentation technique showed that UVPD produced the most amount of unique data, with HCD and ECD close behind, and EID the least (Extended Data Fig. 7c). However, UVPD had significant overlap with EID, which might be a reason for the weak unique proteome coverage result for EID (Extended Data Fig. 7c).
Application of data-independent acquisition in all fragmentation techniques
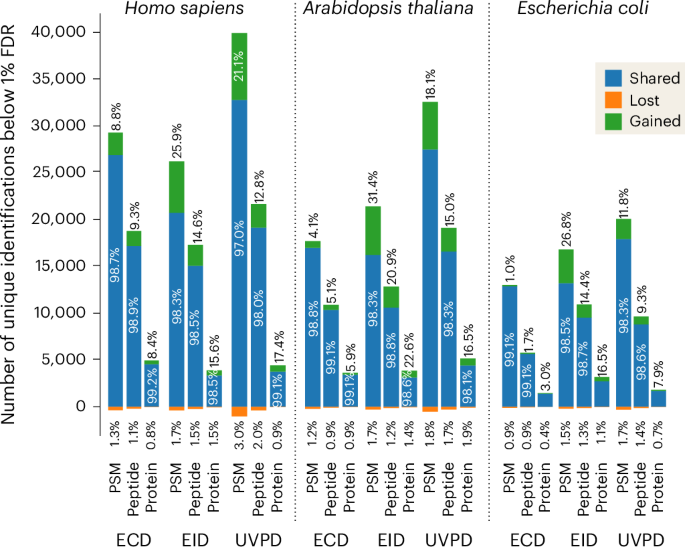
The spectral prediction model created in this work is portable and freely available as ’Prosit_2025_intensity_MultiFrag’ at the Koina model repository47, and can be interfaced from within any software suite. We implemented our model within FragPipe as part of MSBooster29. We reanalyzed the deep proteome data in MSFragger to compare the results with and without MSBooster and found very similar gains to those observed using Oktoberfest at both the PSM and peptide levels (Extended Data Fig. 9). Combined with the optimization of search parameters in FragPipe, we can now perform both data-dependent and data-independent acquisition (DDA and DIA, respectively) analyses (pseudo-DDA through the use of DIA-Umpire) for all activation techniques. The ability to now utilize these activation techniques with DIA approaches led us to create DIA methodologies for the Orbitrap-Omnitrap. The change in ion population, both in terms of ion density and distribution of charge states, required adjustment of the acquisition parameters for each dissociation technique both at the Exploris and Omnitrap level (see Methods). We carried out LC-MS analyses on unfractionated tryptic cell lysate digests from Homo sapiens (Expi293F), Arabidopsis thaliana and Escherichia coli cells. We introduced the last two types of cells to assess the universality of the Prosit model. To optimize duty cycle, we chose to use the ‘normal isolation window’ approach with MS1 range bound to retention time48. MSBooster, using the here-developed Prosit model, increased identification rate at the PSM, peptide and protein levels for all three cell types. The A. thaliana and H. sapiens lysate samples had the largest improvements, trading top position depending on exact context. On average, ECD had the lowest gains across all samples, with the worst result being 1.0%, 1.7% and 3.0% at the three levels for E. coli, while EID demonstrated the largest improvements across all three types of samples, with the best result being 31.4%, 20.9% and 22.6% at the three levels for the A. thaliana sample (Fig. 5).
Number of PSMs, peptides and proteins identified at 1% FDR in the UVPD, EID and ECD DIA data of unfractionated tryptic digests of human, A. thaliana and E. coli proteins. The analysis was performed in the FragPipe platform using the MSFragger search engine with Prosit predictions of fragment ion intensities implemented within the MSBooster module. The numbers of shared, gained and lost identifications correspond to the analysis with MSBooster ’on’ as compared with the results obtained with MSBooster ’off’.